 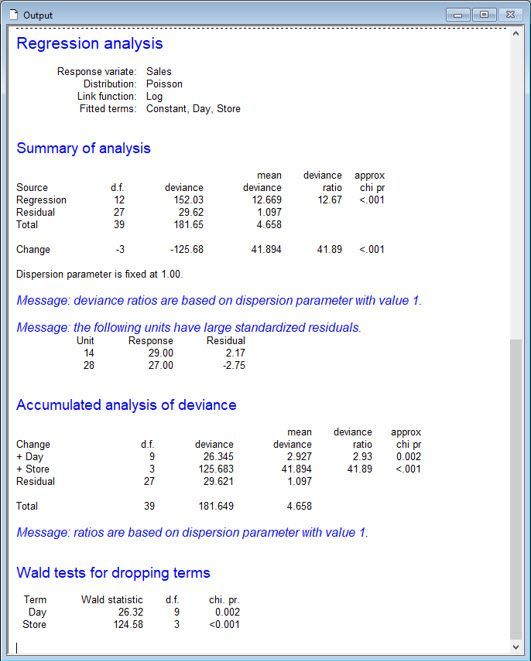
The marker file is recognised by the filename extension.  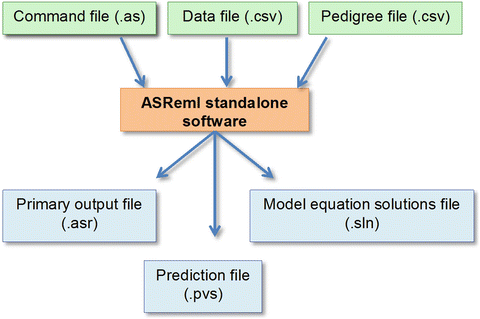
Where p i is the proportion for the minor allele. M is a centred matrix of marker scores (0, 1, 2) One use of the GRM matrix is to fit a Genomic model where G= M M'/s, Large Random Regression (GBLUP from large marker files) The marker map is also copied and applied to this model term It is a group transformation which takes the interval values, and calculates Of marker variables relating to a linkage groupĭefined using !MM which represents additive marker variationĬoded (representing marker states aa, It assumes the argument A is an existing group (if the length is greater than 10, it will be divided by 100 to convertįrom a set of additive marker covariables previously The positions may be given in Morgans or centiMorgans Last marker is taken as the end of the linkage group. The length (right telomere) may be omitted in which case the Linkage group (the position of the right telomere). Telomere position of zero, and an extra value being the length of the Order and coded (backcross) or (F2 design).īe the n marker positions relative to a left This transformation will normally be used on a !G n factor Values to the markers where they have missing values. ASReml Help Genetic Marker operators Setting marker positions !MMAP sĪssigns Haldane map positions ( s) to marker variables and imputes 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
